If conda is installed you should see somehting like the following.Enter conda -V into the terminal command line and press enter.Check conda is installed and in your PATH Anaconda Python distribution installed and accessibleġ.If you have a vanilla Python installation or other Python distribution see virtualenv Outline NOTE: you DO NOT need to install Jupyter in each environment, but you MUST install at least one kernel (for example, ipykernel for Python) in each environment. The conda command is the preferred interface for managing intstallations and virtual environments with the Anaconda Python distribution. Using nbcondakernels you can select which environment to use for every notebook, without leaving Jupyter. Virtual environmets make it easy to cleanly separate different projects and avoid problems with different dependencies and version requiremetns across components. How to set up a virtual environments using conda for the Anaconda Python distributionĪ virtual environment is a named, isolated, working copy of Python that that maintains its own files, directories, and paths so that you can work with specific versions of libraries or Python itself without affecting other Python projects. so file into the directory Pyleoclim_util/pyleoclim/utils where Pyleoclim is installed on your machine.Create virtual environments for python with conda so file will be generated if the compilation is successfulĬopy the. Activating the environment should automatically manage where the linker will look so that the libraries in the environment will be found (this is, in part, what the Conda Forge -compiler packages provide). then run your compilation with this environment activated. 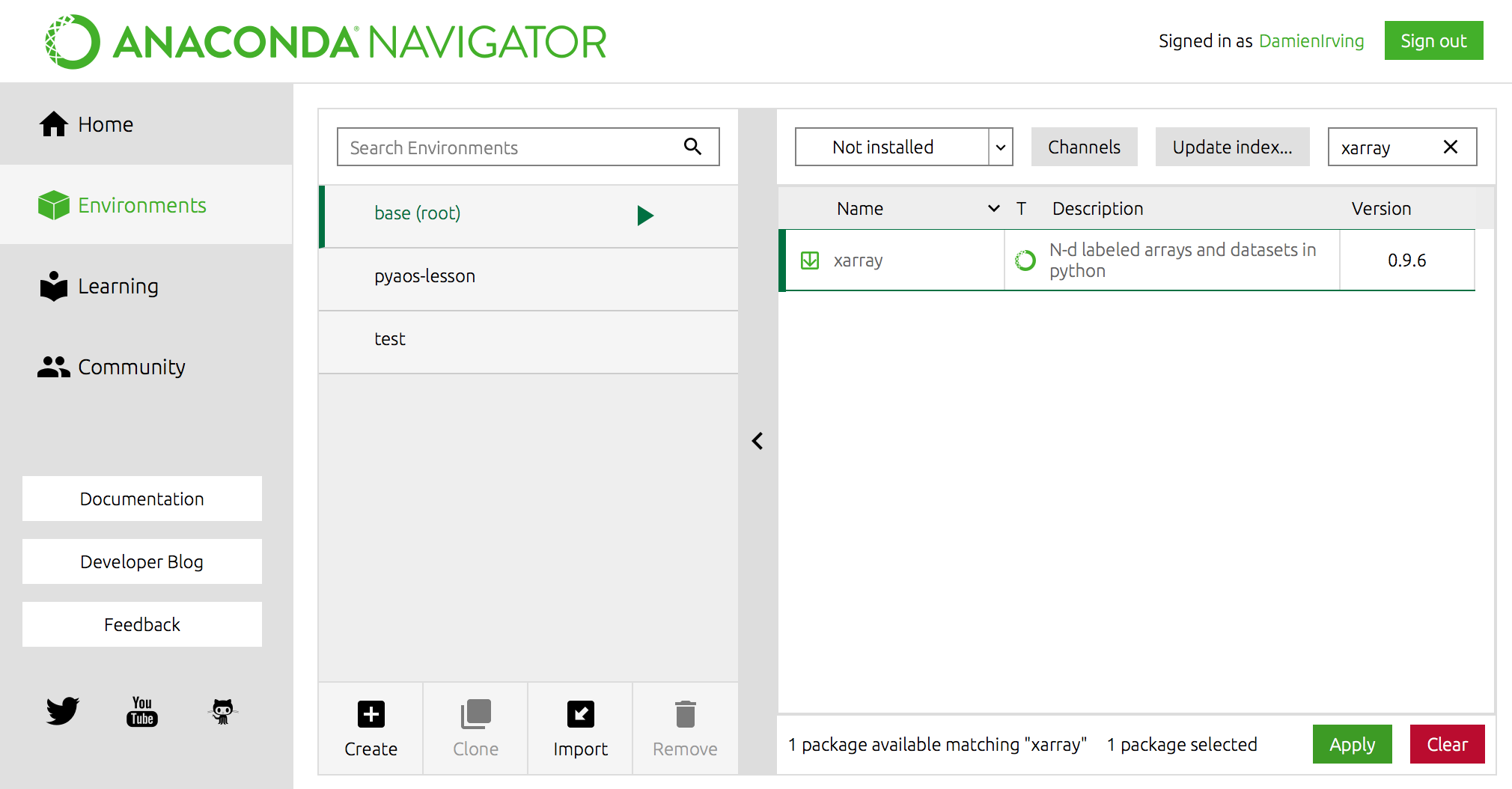
One may also edit the Makefile to use ifort as the compiler to achieve further acceleration just comment out the line for gfortran and use the line for ifort belowĪ. Add packages to Anaconda environment in Python - There are multiple ways by which we can add packages to our existing anaconda environment.Method 1 One comm. conda env create -f fortran-compiler.yaml. Go to the directory Pyleoclim_util/pyleoclim/f2py, and then type make to compile the. gfortran or ifort) is required on your local machine, and the related Fortran source code should be compiled locally following the steps below:ĭownload the source code, either via git clone or just download the. performing it for hundreds of times on timeseries with length longer than 1000 points), in which case we recommend activating the f2py feature to achieve an acceleration of around 50%. However, it could be slow for heavy use (e.g. It is fast enough for lightweight spectral & wavelet analysis tasks, in which case we recommend using the default installation. The default version of WWZ that comes with the installation steps mentioned above is relying on Numba. 
Building from source for the f2py feature of WWZ 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
